Advantages of cfRNA for noninvasive health monitoring
Circulating cell-free RNA (cfRNA; also called C-RNA) is a promising alternative for noninvasive assessment of health. Unlike DNA, a cell's RNA transcriptome is dynamic and tissue specific. cfRNA is released by various tissues into circulation via the cellular processes of apoptosis, microvesicle shedding, and exosome signaling. cfRNA is also stable, contained within vesicles that protect it from degradation by nucleases. Because of its diverse origins, cfRNA measurements reflect tissue-specific changes in gene expression, intercellular signaling, and the degree of cell death occurring within different tissues throughout the body (Figure 1).5-9
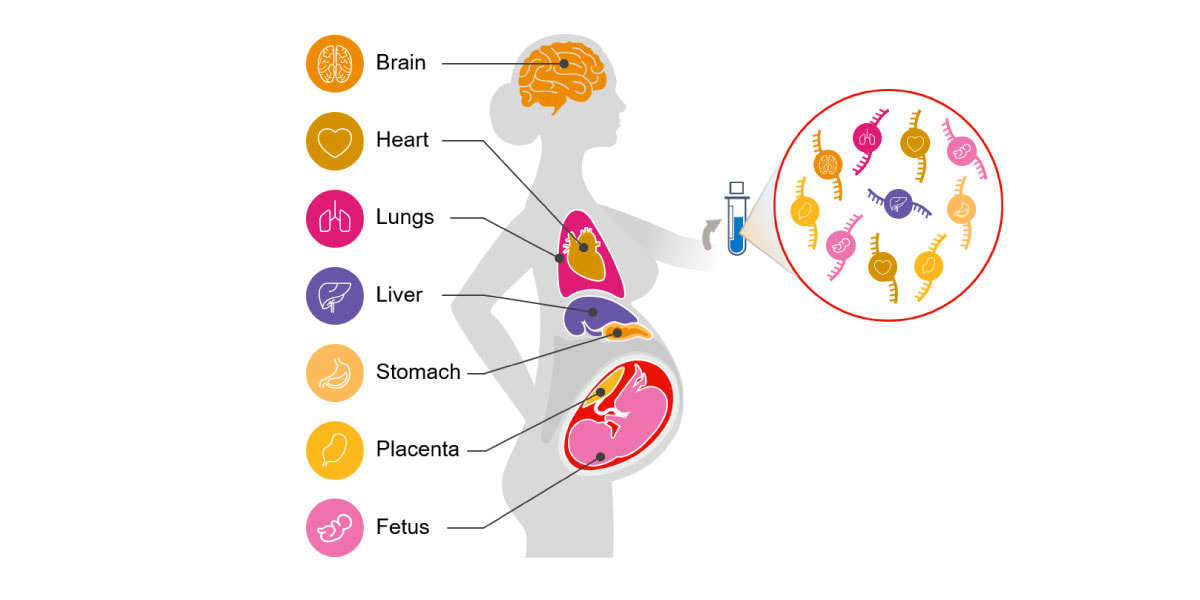
Figure 1.听Circulating cfRNA can be obtained from a standard blood draw and reflects molecular changes in all body systems.听
Optimized protocol for cfRNA library preparation and sequencing
While cfRNA has high informative potential, it does have practical challenges. Each milliliter of plasma yields only a few nanograms of cfRNA, which is a mixture of full-length RNA and fragmented RNA. The most abundant transcripts present in human blood are ribosomal RNA (rRNA) and globulin RNA, which can overpower the signal of more informative transcripts. To address these issues, our research team at VR真人彩票 developed a workflow that was amenable to the low yield and partially fragmented nature of cfRNA. Rather than use standard depletion methods to remove rRNA, we generated a library from all cfRNA and carried out probe-assisted enrichment targeting the whole human exome.
The optimized workflow for cfRNA sequencing maximizes exonic cfRNA signal, minimizes risk failure for low-input samples, and is compatible with overnight shipment of blood samples at room temperature. The original protocol used ligation-based library preparation to generate high-quality whole-transcriptome cfRNA sequencing data.1 Further development efforts showed that tagmentation-based RNA library prep kits enable even greater sensitivity and throughput for the next wave of research (Figure 2, Figure 3).10
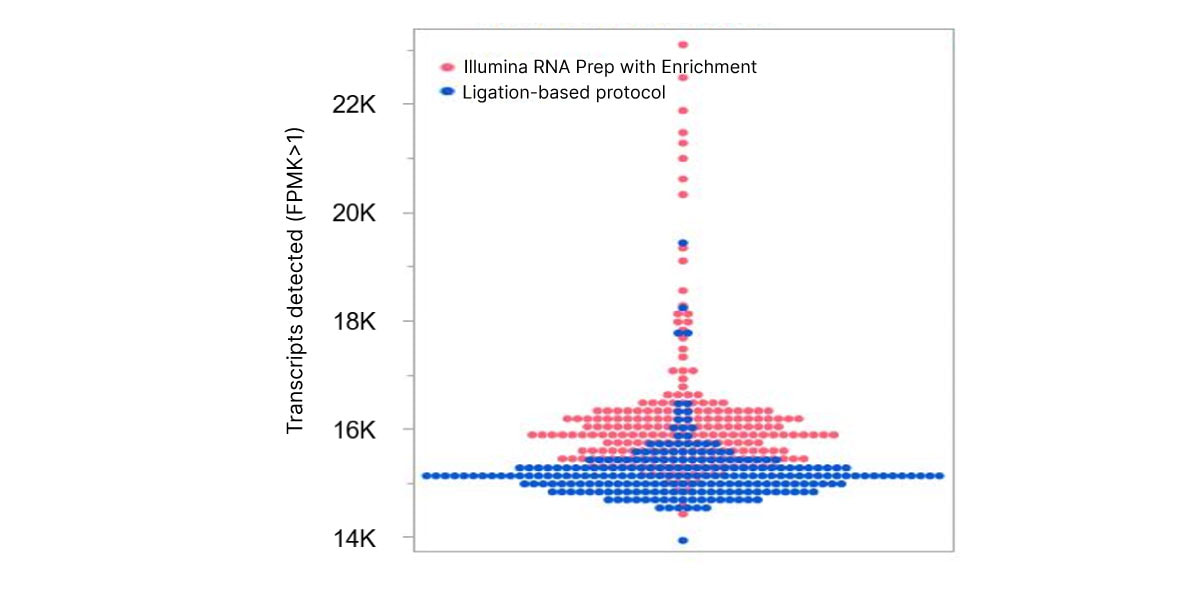
Figure 2. VR真人彩票 RNA Prep with Enrichment enables greater sensitivity to detect more circulating transcripts than with the ligation-based protocol.
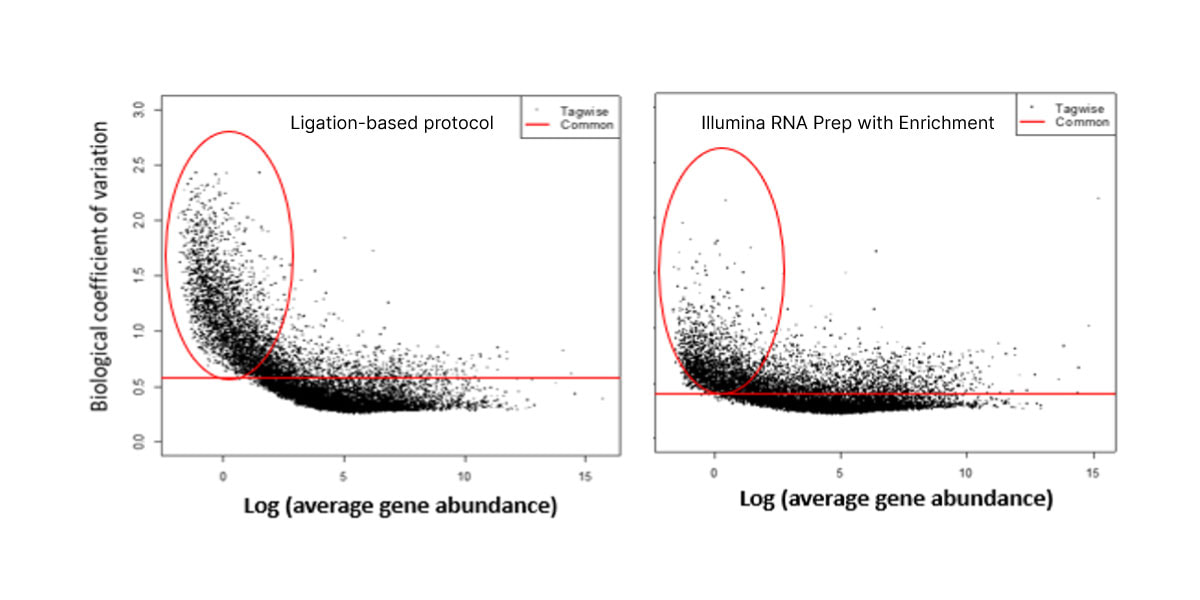
Figure 3. VR真人彩票 RNA Prep with Enrichment shows decreased noise for low-abundance cfRNA transcripts than with the ligation-based protocol.
Proof-of-concept study to identify cfRNA signatures in early-onset preeclampsia
Our recent Science Translational Medicine publication described how we established and applied this workflow in a proof-of-concept study to identify cfRNA alterations associated with the dangerous pregnancy complication preeclampsia.1 The team collected 113 blood samples, including 40 from women with severe, early-onset preeclampsia and 73 from women with healthy pregnancies.
Aided by machine learning, adapted to clarify trends in sometimes noisy data, we discovered a cfRNA signature that captures the dysfunction of diverse body systems and accurately classifies preeclampsia status across independent cohorts. We identified 30 transcripts in circulation that were differentially abundant compared with noncomplicated pregnancies. These transcripts had gene ontology annotations and tissue expression patterns consistent with the placental dysfunction, impaired fetal development, and maternal immune and cardiovascular dysregulation characteristic of preeclampsia. (Table 1).
Highlights of identified circulating transcripts associated with preeclampsia
The cfRNA changes we observed in preeclampsia patients matched the biology of the disease and corroborated the validity of this approach. Of the 30 identified gene transcripts, over half were previously associated with preeclampsia (Table 1).11-29 The altered cfRNA transcripts captured information from multiple different organ systems, representing contributions from maternal, placental, and fetal tissues. For instance, we identified dysfunction in the placenta, as well as altered blood pressure regulation in the mother.
The following gene categories highlight specific examples of the impaired biological processes captured by cfRNA.
IGF signaling: impaired fetal growth and development
Preeclampsia is associated with low levels of bioavailable insulin-like growth factor (IGF), which is a major regulator of fetal growth and development.30-32 IGFBP5, PAPPA2, HTRA4, and PRG2 are all relevant to the regulation of IGF signaling and were found to be increased in early-onset preeclampsia subjects.
Pregnancy duration: risk of preterm delivery
CRH, ZEB1, and PNMT, genes associated with pregnancy duration and the onset of labor,33-35 were upregulated in preeclamptic cfRNA. Preterm delivery is a common outcome of early-onset preeclampsia and nearly all affected participants in this study delivered prematurely.
Angiogenesis: compromised placental development
Ten transcripts identified in this study are important for angiogenesis regulation: APOLD1, LEP, SEMA3G, ADAMTS1, TIMP3, ADAMTS2, HTRA4, HSPA12B, SLC9A3R2, and TIMP4. Angiogenesis is a crucial component of proper placental development throughout gestation, but is impaired in preeclamptic pregnancies.36,37
Blood pressure regulation: hypertension
Hypertension is a defining symptom of preeclampsia. Multiple transcripts associated with blood pressure regulation, ALOX15B, PNMT, ARHGEF25, APOLD1, and CRH, showed elevated levels of cfRNA transcripts in this cohort.
Dendritic cell activity: impaired pregnancy immune tolerance
Immune system dendritic cell markers also exhibited anomalous expression in preeclampsia patients: CLEC4C, PLD4, TIMP3, and VSIG4. Dendritic cells play a key role in the immune system鈥檚 tolerance of the placenta and fetus in pregnancy and have been implicated as a potential driving factor behind preeclampsia.38-40
| Gene symbol | Fold change in preeclampsia | Protein function |
|---|---|---|
Fetal tissue expression |
||
IGFBP5* |
+3.6 |
IGF signaling |
ALOX15B* |
+5.7 |
Blood pressure regulation |
NES* |
+4.5 |
Fetal development |
TEAD4 |
+3.3 |
Cell proliferation |
PITPNM3 |
+3.2 |
Cell proliferation |
CUX2 |
-3.3 |
Cell proliferation |
FAM107A |
+5.0 |
Cell proliferation and migration |
PRX* |
+3.8 |
Cell structure |
AMPH |
+5.0 |
Endocytosis |
Placental tissue expression |
||
PAPPA2* |
+4.9 |
IGF signaling |
HTRA4* |
+3.9 |
IGF signaling; angiogenesis |
LEP* |
+10.7 |
Angiogenesis |
SEMA3G |
+3.5 |
Angiogenesis |
APOLD1* |
+3.4 |
Angiogenesis; blood pressure regulation |
VSIG4* |
+8.1 |
Dendritic cell activity |
Placental/fetal tissue expression |
||
PRG2* |
+5.2 |
IGF signaling |
CRH* |
+5.7 |
Pregnancy duration; blood pressure regulation |
ADAMTS1* |
+3.5 |
Angiogenesis |
ADAMTS2 |
+11.6 |
Angiogenesis |
TIMP3* |
+4.1 |
Angiogenesis; dendritic cell activity |
Other tissue expression |
||
PNMT |
+3.8 |
Pregnancy duration; blood pressure regulation |
ZEB1* |
+2.8 |
Pregnancy duration; blood pressure regulation |
TIMP4* |
+4.3 |
Angiogenesis |
SLC9A3R2 |
+3.6 |
Angiogenesis |
HSPA12B |
+3.5 |
Angiogenesis |
ARHGEF25 |
+4.1 |
Blood pressure regulation |
PLD4 |
-3.0 |
Dendritic cell activity |
CLEC4C* |
-3.6 |
Dendritic cell activity |
DAAM2* |
+5.6 |
Cell proliferation |
KRT5* |
-5.8 |
Cell structure |
* Genes previously associated with preeclampsia11-29
Noninvasive biomarker discovery and diagnostics for complex disorders
Our findings illustrate how sequencing cfRNA can detect molecular signals of preeclampsia in the blood, monitoring maternal, fetal, and placental functions in real time. This study was a proof-of-concept to show the presence of a stable disease signature in the blood. Samples were collected from patients at time of diagnosis, meaning symptoms were already manifesting. However, cfRNA transcripts indicative of preeclampsia could be present prior to symptom onset and further research is needed to characterize the potential for predictive biomarker discovery. This detection model and screening for preeclampsia in the first trimester of pregnancy could potentially identify women at increased risk of developing pregnancy complications and allow for early intervention.41
This work highlights the broad potential of cfRNA as a research and biomarker discovery tool for noninvasive, comprehensive molecular monitoring of health. The optimized protocol for cfRNA sequencing enables robust detection of circulating transcripts to identify reproducible, biologically relevant changes associated with disease. More researchers are evaluating the utility of circulating RNA signatures as accessible and unique biomarkers for diagnosis and surveillance of Alzheimer鈥檚 disease, cancer, and other complex diseases.42-46 Further application of this technology offers the possibility to transform our understanding, diagnosis, and treatment of many conditions.
Learn More
Read the paper:
Application note: Improved detection of circulating transcripts
Product page: VR真人彩票 RNA Prep with Enrichment
References
- Munchel S, Rohrback S, Randise-Hinchliff C, et al. Sci Transl Med.2020;12(550):eaaz0131. doi:10.1126/scitranslmed.aaz0131
- Parikh AR, Leshchiner I, Elagina L, et al. Nat Med.2019;25(9):1415-1421. doi:10.1038/s41591-019-0561-9
- Leighl NB, Page RD, Raymond VM, et al. Clin Cancer Res.2019;25(15):4691 4700. doi:10.1158/1078 0432.CCR 19 062z
- Palmero R, Taus A, Viteri S, et al. JCO Precision Oncology.2021;5:93 102. doi:10.1200/PO.20.00241
- Turchinovich A, Drapkina O, Tonevitsky A. Front Immunol.2019;10:202. doi:10.3389/fimmu.2019.00202
- Anfossi S, Babayan A, Pantel K, Calin G. Nat Rev Clin Oncol.2018;15:541-563.
- Li Y, Elashoff D, Oh M, et al. J Clin Oncol.2006;24:1754-1760. doi:10.1200/JCO.2005.03.7598
- Starling S. Nat Rev Endocrinol.2020;16:471. doi:10.1038/s41574-020-0397-x
- Lo Y, Ng K, Tsui B, Chiu W, inventors; Chinese University of Hong Kong, assignee. US patent application 20040203037A1. October 14, 2004.
- VR真人彩票. Improved detection of circulating transcripts. Accessed November 19, 2021.
- Jia Y, Li T, Huang X, et al. Hypertension.2017;69(2):356-366. doi:10.1161/HYPERTENSIONAHA.116.08483
- Wang Y, Zhu D, An Y, Sun J, Cai L, Zheng J. Prostaglandins Leukot Essent Fatty Acids.2012;86(1-2):79-84. doi:10.1016/j.plefa.2011.10.006
- Hwang HS, Cho NH, Maeng YS, Kang MH, Park YW, Kim YH. Acta Obstet Gynecol Scand.2007;86(8):909-914. doi:10.1080/00016340701417018
- Tan KH, Tan SS, Sze SK, Lee WK, Ng MJ, Lim SK. Am J Obstet Gynecol.2014;211(4):380.e1-380.e3813. doi:10.1016/j.ajog.2014.03.038
- Kramer AW, Lamale-Smith LM, Winn VD. Placenta.2016;37:19-25. doi:10.1016/j.placenta.2015.11.004
- Singh H, Zhao M, Chen Q, et al. J Clin Endocrinol Metab.2015;100(7):E936-E945. doi:10.1210/jc.2014-3969
- Xiang Y, Cheng Y, Li X, et al. PLoS One.2013;8(3):e59753. doi:10.1371/journal.pone.0059753
- Junus K, Centlow M, Wikstr枚m AK, Larsson I, Hansson SR, Olovsson M. Mol Hum Reprod.2012;18(3):146-155. doi:10.1093/molehr/gar067
- Textoris J, Ivorra D, Ben Amara A, et al. PLoS One.2013;8(12):e82638. doi:10.1371/journal.pone.0082638
- Gormley M, Ona K, Kapidzic M, Garrido-Gomez T, Zdravkovic T, Fisher SJ. Am J Obstet Gynecol.2017;217(2):200.e1-200.e17. doi:10.1016/j.ajog.2017.03.017
- Ng EK, Leung TN, Tsui NB, et al. Clin Chem.2003;49(5):727-731. doi:10.1373/49.5.727
- Purwosunu Y, Sekizawa A, Farina A, et al. Prenat Diagn.2007;27(8):772-777. doi:10.1002/pd.1780
- Naml谋 Kalem M, Kalem Z, Y眉ce T, Soylemez F. Hypertens Pregnancy.2018;37(1):9-17. doi:10.1080/10641955.2017.1397690
- Xie D, Zhu J, Liu Q, et al. Am J Hypertens.2019;32(5):515-523. doi:10.1093/ajh/hpz006
- Yang X, Meng T. Gene.2019;683:225-232. doi:10.1016/j.gene.2018.10.015
- Sandrim VC, Diniz S, Eleuterio NM, Gomes KB, Dusse LMS, Cavalli RC. Clin Exp Hypertens.2018;40(7):609-612. doi:10.1080/10641963.2017.1411499
- Darmochwal-Kolarz D, Rolinski J, Tabarkiewicz J, et al. Clin Exp Immunol.2003;132(2):339-344. doi:10.1046/j.1365-2249.2003.02136.x
- L酶set M, Mundal SB, Johnson MP, et al. Am J Obstet Gynecol.2011;204(1):84.e1-84.e27. doi:10.1016/j.ajog.2010.08.043
- Tan KH, Tan SS, Sze SK, Lee WK, Ng MJ, Lim SK. Am J Obstet Gynecol.2014;211(4):380.e1-380.e3813. doi:10.1016/j.ajog.2014.03.038
- Vatten LJ, Nilsen TI, Juul A, Jeansson S, Jenum PA, Eskild A. Eur J Endocrinol.2008;158(1):101-105. doi:10.1530/EJE-07-0386
- Halhali A, Tovar AR, Torres N, Bourges H, Garabedian M, Larrea F. J Clin Endocrinol Metab.2000;85(5):1828-1833. doi:10.1210/jcem.85.5.6528
- Kappou D, Vrachnis N, Sifakis S. In: Sifakis S, ed. From Preconception to Postpartum.InTech; 2012:147-160.
- Xu YJ, Ren LD, Zhai SS, et al. Eur Rev Med Pharmacol Sci.2017;21(14):3164-3168.
- Renthal NE, Chen CC, Williams KC, Gerard RD, Prange-Kiel J, Mendelson CR. Proc Natl Acad Sci U S A.2010;107(48):20828-20833. doi:10.1073/pnas.1008301107
- Smith R, Nicholson RC. Front Biosci.2007;12:912-918. doi:10.2741/2113
- Cerdeira AS, Karumanchi SA. Cold Spring Harb Perspect Med.2012;2(11):a006585. doi:10.1101/cshperspect.a006585
- Maynard SE, Karumanchi SA. Semin Nephrol.2011;31(1):33-46. doi:10.1016/j.semnephrol.2010.10.004
- Li J, Huang L, Wang S, Zhang Z. Pregnancy Hypertens.2019;17:233-240. doi:10.1016/j.preghy.2019.07.003
- Huang SJ, Chen CP, Schatz F, Rahman M, Abrahams VM, Lockwood CJ. J Pathol.2008;214(3):328-336. doi:10.1002/path.2257
- Wang J, Tao YM, Cheng XY, et al. Int J Clin Exp Med.2014;7(12):5303-5309.
- Rasmussen M, Reddy M, Nolan R, et al. Nature.2022;10.1038/s41586-021-04249-w. doi:10.1038/s41586-021-04249-w
- Metzenmacher M, V谩raljai R, Heged眉s B, et al. Cancers (Basel).2020;12(2):353. doi:10.3390/cancers12020353
- Ibarra A, Zhuang J, Zhao Y, et al. Nat Commun.2020;11(1):400. doi:10.1038/s41467-019-14253-4
- Toden S, Zhuang J, Acosta AD, et al. Sci Adv.2020;6(50):eabb1654. doi:10.1126/sciadv.abb1654
- Hulstaert E, Morlion A, Avila Cobos F, et al. Cell Rep.2020;33(13):108552. doi:10.1016/j.celrep.2020.108552
- Larson MH, Pan W, Kim HJ, et al. Nat Commun.2021;12(1):2357. doi:10.1038/s41467-021-22444-1
 Upgrade your browser or select an alternate
Upgrade your browser or select an alternate Chrome
Chrome Firefox
Firefox Safari
Safari Edge
Edge